A molecular view on protein retrotranslocation
Quality control in eukaryotic cells
Researchers at Harvard Medical School (USA), New York University School of Medicine (USA) and the Max Planck Institute of Biophysics developed a model of how misfolded proteins are recognised and retrotranslocated by the Hrd1 ubiquitin ligase complex as part of the ER-associated protein degradation pathway (ERAD) based on data from cryo-electron microscopy, biochemical experiments and molecular simulations.
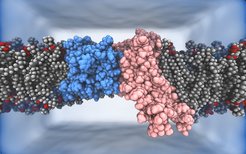
Snapshot of the molecular simulation of the membrane components of the Hrd1 ubiquitin ligase complex, Hrd1 (pink) and Der1 (blue), in a lipid bilayer.
Whenever newly produced proteins are misfolded in the endoplasmic reticulum (ER) they have to be degraded to maintain homeostasis. For this they must be retrotranslocated into the cytosol and polyubiquitinated to be ultimately degraded by the proteasome. This pathway is called ER-associated protein degradation, ERAD. The details of this process are largely unclear and a major unsolved question is how misfolded peptides can cross the ER membrane, which usually constitutes a barrier to proteins and other molecules.
To shed light on this process a research team at Harvard University and the New York University School of Medicine under the lead of Tom Rapoport succeeded in solving the structure of the active Hrd1 complex from S. cerevisiae using cryo-electron microscopy. Then they established the functional importance of this complex both with biochemical and in vivo experiments. Marc Siggel from Max Planck Institute of Biophysics, Department of Theoretical Biophysics, led by Gerhard Hummer, examined this question by molecular simulations to gain insights into the dynamics of this complex and its membrane-based components Hrd1 and Der1 at an atomistic level. Computer simulations revealed details about the behavior of the complex in its membrane environment which are otherwise difficult to obtain solely by experimental methods.
With these combined data a molecular model of the mechanism of translocation was established: The substrate can be moved through two half channels formed by Hrd1 and Der1 which are juxtaposed in a thinned and distorted membrane region. This altered membrane environment facilitates movement of hydrophilic substrates across the membrane and might be a general concept used by many protein translocation systems.
Misfolded proteins are prevented from travelling to their destination, extracted from the ER and delivered to the proteasomal degradation machinery in the cytoplasm. This work provides an important cornerstone for understanding the degradation pathway at the molecular level and how this complex macromolecular machinery functions.
