Structure of bd Oxidase
On the trail of a protein essential for survival in mycobacteria
Researchers from the University of Otago, the Goethe University Frankfurt and the Max Planck Institute of Biophysics solved the cryo-EM structure of the cytochrome bd oxidase from Mycobacterium tuberculosis at a resolution of 2.5 Å. In conjunction with atomistic molecular dynamics simulations a previously unknown MK-9-binding site was discovered, as well as a unique disulfide bond within the Q-loop domain that defines an inactive conformation of the canonical quinol oxidation site in Actinobacteria.
In 1882, Robert Koch identified Mycobacterium tuberculosis (M. tb) as the primary cause of tuberculosis (TB) - a ground-breaking discovery honored with the Nobel Prize in Physiology or Medicine. Today, over a hundred years later, TB is still the major leading cause of death by an infectious disease worldwide. TB kills approximately 5000 people a day and the total count of TB deaths before anti-TB drugs were discovered tallies up to 1 billion people. The greatest challenge fighting TB in the antibiotic era is the ever-growing emergence of multi and extensive drug resistant strains. In order to explore innovative paths for identifying novel drug targets, researchers of the Departments of Molecular Membrane Biology and Theoretical Biophysics at the Max Planck Institute of Biophysics headed by Hartmut Michel and Gerhard Hummer together with researchers of the University of Otago (New Zealand) have elucidated the molecular structure of the cytochrome bd oxidase from M. tuberculosis.
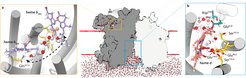
This enzyme is embedded in the bacterial cytoplasmic membrane. It is a crucial component of the adaptive respiratory chain of M. tb and confers mycobacteria with the ability to maintain respiration in the human host organism where free oxygen is very limited. The cytochrome bd oxidase is further critically important for the transition from acute TB infections to dormant states of disease.
This study reveals a unique molecular framework differing from related enzymes and forms the basis for the design and development of inhibitory drugs that can act on M. tb. Treatment of TB via this novel mechanism of action will help to reduce treatment times from up to 12 months to a few weeks.
Furthermore, inhibition of crucial physiological processes like respiration reduces the probability for the emergence of escape mutations, especially important when drugs targeting the respiratory chain are combined.












